2. Key Laboratory of Stress Physiology and Ecology in Cold and Arid Regions, Northwest Institute of Eco-Environment and Resources, Chinese Academy of Sciences, Lanzhou, Gansu 730000, China;
3. Urat Desert-grassland Research Station, Northwest Institute of Eco-Environment and Resources, Chinese Academy of Sciences, Lanzhou, Gansu 730000, China
Artemisia halodendron Turcz. ex Bess. (Asteraceae, Anthemideae, Subgen. Dracunculus) is one of the most common semishrub species in the Horqin Sandy Land in Northeast China. It is important for vegetation rehabilitation in Horqin Sandy Land because of its high ecological value, including that (1) it is the key species of the plant communities and landscapes studies in Horqin Sandy Land (Li, 1991); (2) it plays a key role in the vegetation-restoration process due to its high drought tolerance, anti-wind erosion properties, and sand-burial resistance (Dong et al., 2000 ; Li et al., 2002 ; Zhao et al., 2006 ). Previous studies on A. halodendron focused on aspects of population-distribution patterns (Chao et al., 1999 ; Cao et al., 2008 ), biomass allocation (Li et al., 2005 ), breeding distribution (Li et al., 2005 ), morphological characteristics and physiological adaptations (Zhou et al., 1999 ), root longevity (Huang et al., 2009 ), genetic diversity (Huang et al., 2011 , 2014), and establishment (Li et al., 2002 ) in Horqin Sandy Land. However, systematic comparison of genetic relationships of A. halodendron among populations from different hydrothermal regions has not yet been reported.
Horqin Sandy Land is located in the agropastoral transitional zone between the Inner Mongolian Plateau and the Northeast Plains (42°41′N–45°45′N, 118°35′E–123°30′E) and is one of the four largest sandy areas in northern China; it covers an area of approximately 139,300 km2, of which up to 71,884 km2 is desertified sandy land (Wang, 2003; Zhao et al., 2003 ). Landscape in this area is characterized by sand dunes that alternate with gently undulating lowland areas (Li et al., 2005 ). This area belongs to the continental semi-arid monsoon climate and is in the temperate zone, with a mean annual temperature (AMT) of 3–7 °C and mean annual rainfall (AP) of 350–500 mm (Zhao et al., 2003 ). Over recent decades, this region has undergone severe desertification (Li et al., 2000 , 2004) and has displayed the northern-moving phenomenon of the interlocked agropasturing area of North China in the most recent hundred years (Zhao et al., 2000 , 2002).
In the present study, we used chloroplast DNA (cpDNA) trnL–F to examine the genetic diversity ofA. halodendron. We specifically aimed to address the following questions: (1) What is the level of nucleotide diversity in A. halodendron from different hydrothermal regions? (2) How are the identified haplotypes distributed within and among populations? In particular, is there subdivision in the different hydrothermal-level populations (according to hydrothermal synthesis index)? We attempted to interpret the results to provide baseline genetic information pertinent to the restoration and management of degraded ecosystems in arid and semi-arid areas.
2 Materials and methods 2.1 SamplingA total of 243 individuals were sampled from 10 natural A. halodendron populations (Tables 1 and 2). Populations 1–6 were in the low-hydrothermal-synthesis index region (average 24.97), while populations 7–10 were in the high-hydrothermal-synthesis index region (average 31.98) (Tables 1 and 2). The equation of hydrothermal synthesis index was
|
|
Table 1 The average monthly rainfall in 10 years of 10 populations of Artermisia halodendron in Horqin Sandy Land |
|
|
Table 2 The monthly mean temperature in 10 years of 10 populations for Artermisia halodendron in Horqin Sandy Land |
Total genomic DNA was extracted using AxyPrep Genomic DNA Mini Kits (Axygen Inc., Beijing, China) following the manufacturer's instructions. DNA quality was checked on a 1.0% agarose gel. Several pairs of cpDNA primers designed by Hamilton (1999), Taberlet et al. (1991) , and Sang et al. (1997) were used in the initial screening. Two pairs of primers, trnL (5'-CGGAATTGGTAGACGCTACG-3') and trnF (5'-ATTTGAACTGGTGACACGAG-3') (Sang et al., 1997 ), identified sequence variations in the sampled individuals and therefore were used for all remaining individuals. Polymerase chain reaction (PCR) was performed in a 25-μL reaction volume, containing 40 ng of genomic DNA, 1.0 U of Taq polymerase (Axygen Inc., Beijing, China), 3 mmol/L MgCl2, 500 μmol/L each dNTP, 20 mmol/L Tris-HCl (pH 8.3), 100 mmol/L KCl, and 0.3 μmol/L primer. The amplification condition was an initial denaturation step at 94 °C for 3 min, followed by 30 cycles of 30 s at 94 °C, 30 s at 55 °C, 1 min at 72 °C, and a final 5-min extension step at 72 °C. The PCR products were determined by 1.0% agarose gel electrophoresis. The amplification products were purified using an AxyPrep PCR Purification Kit, following the manufacturer's protocol (Axygen Inc., Beijing, China). Purified DNA was sequenced by the MEIJI sequencing company in Shanghai, China, applying the PCR-primers as sequencing primers.
2.3 Data analysesDNA sequences were aligned using the CLUSTAL X program (Thompson et al., 1997 ), with subsequent manual adjustments in MEGA4 (Tamura et al., 2007 ). A matrix of combined sequences was constructed for the 243 individuals that we examined, and different cpDNA sequences were identified as haplotypes.
Basic population genetic parameters were estimated for three groups of populations: the low-hydrothermal-level region (populations 1–6); the high-hydrothermal-level region (populations 7–10); and finally, all populations. All parameters were calculated with DNASP 5.10.01 (Librado and Rozas, 2009), including the number of segregating sites (S), the number of haplotypes (Nh), the haplotype diversity (Hd), the average number of nucleotide differences per site between two sequences in a sample, π (Nei and Li, 1979; Nei, 1987), and the average number of pairwise nucleotide differences (k).
Phylogenetic analyses of cpDNA haplotypes were performed with maximum parsimony (MP), using PAUP version 4.0 (Swofford, 2002). Heuristic search was implemented with 100 random additional sequence replicates, tree-bisection-reconnection (TBR) branch swapping, MULPARS option, and ACCTRAN optimization. To evaluate the relative robustness of the clades found in the most parsimonious tree, bootstrap analysis was conducted using 1,000 replicates with a simple taxon addition. Genetic differentiation among populations at the three different sampling levels was estimated by pairwise FST values (Wright, 1951). AMOVA was performed to analyze the source of variation among populations, using Arlequin 3.0 (Excoffier et al., 2005 ) with 1,000 replicates of bootstrap.
3 Results 3.1 Haplotype distribution and genetic diversitySequence data were obtained for one loci from on average 101, 142, and 243 individuals from the low-hydrothermal-level region, the high-hydrothermal-level region, and the entire species' range, respectively. The length of the aligned trnL–trnF DNA sequences (including trnL and the trnF spacer region) ranged between 849 and 863 bp with two insertions. The analysis of cpDNA variation identified seven haplotypes (HapA-HapG) (Table 4). Haplotype C was the most abundant, occurring in three populations, followed by haplotypes A, B, D, and G, which occurred in two populations; and the remaining haplotypes were found in only a single population (Table 3 and Figure 1). We confirmed the division of the native range into two areas using hydrothermal synthesis index data (Table 3). Six populations in the low-hydrothermal-level region of the species were dominated by four different haplotypes. The other four populations in the high-hydrothermal-level region of the species were dominated by three different haplotypes (Table 3 and Figure 1).
|
|
Table 3 Origin of materials and number of samples for 10 populations of Artemisia halodendron from the Horqin Sandy Land |

|
Figure 1 Geographic distribution of the seven haplotypes found on 243 individuals for trnL–F observed inArtemisia halodendron of the Horqin Sandy Land. Pie charts indicate the frequency of haplotypes within each population, and unique alleles are indicated by different colours. NDRS indicates Naiman Desertification Research Station, China Academy of Sciences. Base map data produced in 2000 |
|
|
Table 4 Variable sites of the aligned sequences of trnL-F in seven haplotypes of Artemisia halodendron in the Horqin Sandy Land |
We identified a total of two, three, and three segregating sites for the low-hydrothermal-level region, the high-hydrothermal-level region, and all populations, respectively. Summary statistics of sequence variation are given in Table 5. Overall haplotype diversity (Hd) and nucleotide sequence (π) diversity for A. halodendron were 0.706±0.001 and 0.0013±0.0001, respectively. At the regional level, haplotype diversity and nucleotide diversity between two regions varied between 0.318 (low-hydrothermal-level region) and 0.671 (high-hydrothermal-level region), and between 0.0006 (low-hydrothermal-level region) and 0.0015 (high-hydrothermal-level region), respectively. The population from the high-hydrothermal-level region had higher haplotype diversity and nucleotide diversity than the population from the low-hydrothermal-level region.
|
|
Table 5 Nucleotide variation and haplotype diversity at trnL-F in 10 populations of Artemisia halodendron from the low-hydrothermal-level region (populations 1–6), high-hydrothermal-level region (populations 7–10), and entire species' range (all populations) in the Horqin Sandy Land |
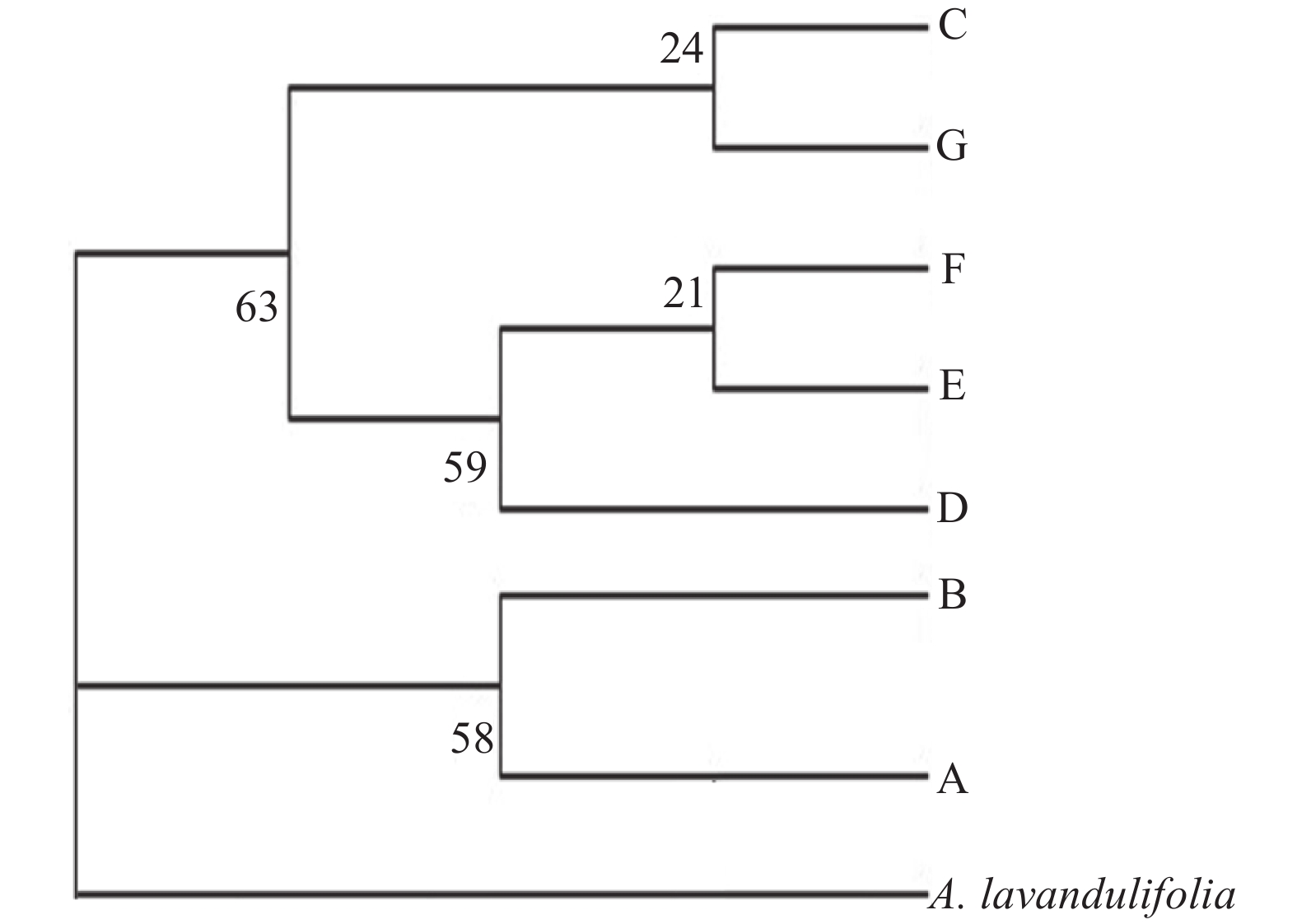
Maximum parsimony analysis resulted in a single tree (length=14 and consistency index =0.5834). Two clades with low bootstrap support were identified: one consisting of the Hap C, G, F, E, and D, with four haplotypes (Hap C, G, F, and Hap D) distributed in the low-hydrothermal-level region and one haplotype (Hap E) in the high-hydrothermal-level region; and the other consisting of the rest of Hap A and Hap B, both occurring in the high-hydrothermal-level region (Figure 2).
|
Figure 2 Phylogenetic relationships of the identified haplotypes of Artemisia halodendron in Horqin Sandy Land |
The analysis of molecular variance (AMOVA) showed that around 17.54% of the variation was attributed to the low-/high-hydrothermal-level regions differentiation, and between- populations variation accounted for just over one-half (55%) of the total variation, indicating very strong differentiation and little gene flow between regions and populations (Table 6). Within the low- and high-hydrothermal-level regions, between-populations differentiation was respectively 81.52% and 93.77% (Table 6). Overall, these results strongly indicate that haplotypes are geographically structured across the species' distribution range. FST values were 0.728, 0.815, and 0.938 at the species level, low-hydrothermal-level region, and high-hydrothermal-level region, respectively.
|
|
Table 6 Results of analyses of molecular variance (AMOVAs) of haplotype frequencies for populations and regional populations of Artemisia halodendron in Horqin Sandy Land |
In this study, we examined nucleotide variation at two nuclear loci of A. halodendron sampled from the Horqin Sandy Land. We confirmed the division of the native range into two areas, using hydrothermal synthesis index data (Table 3). The analysis of cpDNA variation identified seven haplotypes (Table 4 and Figure 2). Six populations in the low-hydrothermal-level region of the species were dominated by four different haplotypes. The other four populations in the high-hydrothermal-level region of the species were dominated by three different haplotypes (Table 3 and Figure 2).
Many studies have demonstrated that endemic species tend to possess high levels of genetic diversity (Wang et al., 2010 ; Ge et al., 2011 ). Compared with similar studies (Wang et al., 2011 ), the level of haplotype diversity (Hd=0.706) and the nucleotide diversity (π=0.0013) within the whole populations of A. halodendron were somewhat lower than that of the endemic plants in northern of China. Such a low Hd value and π value indicated that this species was very adapted to the sandy land environment, and this character might have made it become the dominant and constructive species in Horqin Sandy Land. At the same time, the populations from the low-hydrothermal-level region exhibited lower haplotype diversity and more limited nucleotide diversity, and were a subset of that observed in the populations from the the high-hydrothermal-level region. These results lend support to the scenario described by Huang et al. (2011) using ISSR markers and Huang et al. (2013) using psbA-trnH. This fact might be due to environmental differences (Li et al., 2009 ; Bao et al., 2010 ; Liu et al., 2010 ; Li and Wu, 2011). The low-hydrothermal-level region is a region of severe desertification (Zhao et al., 2000 ; Bao et al., 2010 ). In the process of ecological restoration, species increasing adapt to the local environment, which, in turn, reduces the genetic diversity (Wang et al., 2009 ; Huang et al., 2011 ).
Phylogenetic relationships analysis of the cpDNA sequences collected from A. halodendron in the different hydrothermal-level regions in Horqin Sandy Land did not entirely cluster according to populations or hydrothermal-level regions. This result is similar to that of some other studies (Chen et al., 2009 ; Hu et al., 2010 ; Zhou et al., 2010 ; Zhang et al., 2015 ). The MP tree indicated that the seven haplotypes formed two clades: the haplotypes in clade I all came from the high-hydrothermal-level region (63% bootstrap support); the haplotypes (except Hap E) in clade II came from the low-hydrothermal-level region (58% bootstrap support). This cluster process was closely related to the hydrothermal gradients in Horqin Sandy Land. The results of phylogenetic relationships analysis show that hydrothermal conditions change determines the increase of the haplotypes distribution difference.
The spatial analyses of genetic variation in A. halodendron in the different hydrothermal-level regions in Horqin Sandy Land indicate that the between-populations genetic differentiations are high within the low-, the high-hydrothermal-level region, and the total distribution range (Table 6). Our data revealed significant genetic differences among populations in different hydrothermal level regions. The increase in population genetic differentiation might be related to strong human disturbance, high rates of habitat fragmentation, and decreasing population size in Horqin Sandy Land (Zhao et al., 2000 ; Wang et al., 2010 ). It is generally accepted that genetic differentiation would have increased among the native populations due to the changed environment (Wang et al., 2009 ).
In conclusion, there are significant genetic differences between A. halodendron populations from different hydrothermal-level regions. This information about genetic variation has important implications for restoring and managing the degraded ecosystems in arid and semi-arid areas. It is particularly important to understand the genetic variation of extant populations over the entire range of the species' distribution. Additional research is needed to determine levels of genetic variation in A. halodendron throughout its entire distributional range.
Acknowledgments:The authors thank all the members of Naiman Desertification Research Station, China Academy of Sciences (CAS), for their help in the field work. We acknowledge the China Meteorological Administration (Beijing, China) for help on the meteorological data information support. This study was financially supported by research projects 2016YFC0500907, 2017FY100205, 41201561, Y551821001, and 145RJYA269.
Bailey HP, 1979. Semi-arid climates: their definition and distribution. In: Hall AE, Cannell GH, Lawton HW (eds.). Agriculture in Semi-Arid Environments Berlin, Heidelberg: Springer, pp. 73–97. DOI: 10.1007/978-3-642-67328-3_3.
|
Bao HJ, Guo J, Yan L. 2010. Study on ecological footprint of the human activity intensity in Horqin sandy—A case of Naiman Banner. Journal of Arid Land Resources and Environment, 24(2): 126-131. DOI:10.13448/j.cnki.jalre.2010.02.012 |
Cao YN, Shi LS, Han S, et al. 2008. Point pattern analysis for a population of Artemisia halodendron in a Kerqin Sandlot
. Chinese Bulletin of Botany, 25(4): 437-442. DOI:10.3969/j.issn.1674-3466.2008.04.007 |
Chao LM, Piao SJ, Zhi RN, et al. 1999. The distribution patterns of Artemisia halodendron in different sandland types
. Journal of Desert Research, 19(S1): 45-48. |
Chen FJ, Wang AL, Chen KM, et al. 2009. Genetic diversity and population structure of the endangered and medically important Rheum tanguticum (Polygonaceae) revealed by SSR Markers
. Biochemical Systematics and Ecology, 37(5): 613-621. DOI:10.1016/j.bse.2009.08.004 |
Dong ZB, Wang XM, Liu LY. 2000. Wind erosion in arid and semiarid China: an overview. Journal of Desert Research, 20(2): 134-139. DOI:10.3321/j.issn:1000-694X.2000.02.007 |
Excoffier L, Laval G, Schneider S. 2005. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online, 1: 47-50. |
Ge XJ, Hwang CC, Liu ZH, et al. 2011. Conservation genetics and phylogeography of endangered and endemic shrub Tetraena mongolica (Zygophyllaceae) in Inner Mongolia, China
. BMC Genetics, 12: 1. DOI:10.1186/1471-2156-12-1 |
Hamilton MB. 1999. Four primer pairs for the amplification of chloroplast intergenic regions with intraspecific variation. Molecular Ecology, 8(3): 521-523. |
Hu YP, Wang L, Xie XL, et al. 2010. Genetic diversity of wild populations of Rheum tanguticum endemic to China as revealed by ISSR analysis
. Biochemical Systematics and Ecology, 38(3): 264-274. DOI:10.1016/j.bse.2010.01.006 |
Huang G, Zhao XY, Huang YX, et al. 2009. The root longevity of Artemisia halodendron inhabiting two sandy land habitats
. Chinese Journal of Plant Ecology, 33(4): 755-763. DOI:10.3773/j.issn.1005-264x.2009.04.014 |
Huang WD, Zhao XY, Zhao X, et al. 2011. A combined approach using ISSR and ITS analysis for the characterization of Artemisia halodendron from Horqin sandy land, northern China
. Biochemical Systematics and Ecology, 39(4–6): 346-351. DOI:10.1016/j.bse.2011.04.011 |
Huang WD, Zhao XY, Zhao X, et al. 2013. Genetic diversity in Artemisia halodendron (Asteraceae) based on chloroplast DNA psbA-trnH region from different hydrothermal conditions in Horqin sandy land, northern China
. Plant Systematics and Evolution, 299: 107-113. DOI:10.1007/s00606-012-0707-4. |
Huang WD, Zhao XY, Zhao X, et al. 2014. Relationship between the genetic diversity of Artemisia halodendron and climatic factors
. Acta Oecologica, 55: 97-103. DOI:10.1016/j.actao.2013.12.005 |
Li FR, Zhang AS, Duan SS, et al. 2005. Patterns of reproductive allocation in Artemisia halodendron inhabiting two contrasting habitats
. Acta Oecologica, 28(1): 57-64. DOI:10.1016/j.actao.2005.02.005 |
Li FR, Zhao LY, Zhang H, et al. 2004. Wind erosion and airborne dust deposition in farmland during spring in the Horqin Sandy Land of eastern Inner Mongolia, China. Soil and Tillage Research, 75(2): 121-130. DOI:10.1016/j.still.2003.08.001 |
Li J. 1991. The distribution of Artemisia halodendron and its status in the natural vegetation succession
. Journal of Desert Research, 11(2): 55-60. |
Li SG, Harazono Y, Oikawa T, et al. 2000. Grassland desertification by grazing and the resulting micrometeorological changes in Inner Mongolia. Agricultural and Forest Meteorology, 102(2–3): 125-137. DOI:10.1016/S0168-1923(00)00101-5 |
Li SG, Harazono Y, Zhao HL, et al. 2002. Micrometeorological changes following establishment of artificially established artemisia vegetation on desertified sandy land in the Horqin Sandy Land, China and their implication on regional environmental change. Journal of Arid Environments, 52(1): 101-119. DOI:10.1006/jare.2001.0983 |
Li Y. 2011. Evaluation on forestry suitability in the Horqin Sandy Land—A case study in Horqinzuoyihouqi Banner. Research of Soil and Water Conservation, 18(6): 236-239, 244. |
Li YQ, Zhao HL, Li YL, et al. 2009. Soil nitrogen mineralization and nitrification in different habitats, Horqin sandy land. Journal of Desert Research, 29(3): 438-444. |
Librado P, Rozas J. 2009. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25(11): 1451-1452. DOI:10.1093/bioinformatics/btp187 |
Liu Y, Zhang DY, Yang HL, et al. 2010. Fine-scale genetic structure of Eremosparton songoricum and implication for conservation
. Journal of Arid Land, 2(1): 26-32. DOI:10.3724/SP.J.1227.201000026 |
Nei M, 1987. Molecular Evolutionary Genetics. New York: Columbia University Press.
|
Nei M, Li WH. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America, 76(10): 5269-5273. |
Sang T, Crawford DJ, Stuessy TF. 1997. Chloroplast DNA phylogeny, reticulate evolution, and biogeography of Paeonia (Paeoniaceae)
. American Journal of Botany, 84(8): 1120-1136. DOI:10.2307/2446155 |
Swofford DL, 2002. PAUP*: Phylogenetic analysis using parsimony (*and other methods), version 4. 0b10. Sunderland, Massachusetts, USA: Sinauer Associates.
|
Taberlet P, Gielly L, Pautou G, et al. 1991. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Molecular Biology, 17(5): 1105-1109. DOI:10.1007/BF00037152 |
Tamura K, Dudley J, Nei M, et al. 2007. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24(8): 1596-1599. DOI:10.1093/molbev/msm092 |
Thompson JD, Plewniak F, Poch O. 1997. A comprehensive comparison of multiple sequence alignment programs. Nucleic Acids Research, 27(13): 2682-2690. DOI:10.1093/nar/27.13.2682 |
Wang CH, Li SF, Fu CZ, et al. 2009. Molecular genetic structure and evolution in native and colonized populations of the Chinese mitten crab, Eriocheir sinensis
. Biological Invasions, 11(2): 389-399. DOI:10.1007/s10530-008-9256-8 |
Wang JF, Pan YZ, Gong X, et al. 2011. Chloroplast DNA variation and phylogeography of Ligularia tongolensis (Asteraceae), a species endemic to the Hengduan Mountains region of China
. Journal of Systematics and Evolution, 49(2): 108-119. DOI:10.1111/j.1759-6831.2011.00117.x |
Wang T, 2003. Desert and Desertification in China. Shijiazhuang: Hebei Science and Technology Press.
|
Wang TJ, Li WQ, Zhang SY, et al. 2010. Genetic diversity and differentiation of five natural populations of Artemisia halodendron
. Scientia Silvae Sinicae, 46(12): 171-175. |
Wright S. 1951. The genetical structure of populations. Annals of Human Genetics, 15(1): 323-354. DOI:10.1111/j.1469-1809.1949.tb02451.x |
Zhang HX, Zhang ML, Wang LN. 2015. Genetic structure and historical demography of Malus sieversii in the Yili Valley and the western mountains of the Junggar Basin, Xinjiang, China
. Journal of Arid Land, 7(2): 264-271. DOI:10.1007/s40333-014-0044-2 |
Zhao HL, Su YZ, Zhou RL. 2006. Restoration mechanism of degraded vegetation in sandy areas of northern China. Journal of Desert Research, 26(3): 323-328. DOI:10.3321/j.issn:1000-694X.2006.03.001 |
Zhao HL, Zhao XY, Zhang TH. 2000. Causes, processes and countermeasures of desertification in the interlocked agro-pasturing area of North China. Journal of Desert Research, 20(S1): 22-28. |
Zhao HL, Zhao XY, Zhang TH, et al. 2002. Boundary line on agro-pasture zigzag zone in North China and its problems on eco-environment. Advance in Earth Sciences, 17(5): 739-747. DOI:10.3321/j.issn:1001-8166.2002.05.017 |
Zhao HL, Zhao XY, Zhang TH, et al., 2003. Desertification Process and Its Recovery Mechanism in Horqin Sandy Land. Beijing: China Ocean Press.
|
Zhou GY, Yang LC, Li CL, et al. 2010. Genetic diversity in endangered Notopterygium forbesii Boissieu based on intraspecies sequence variation of chloroplast DNA and implications for conservation
. Biochemical Systematics and Ecology, 38(5): 911-916. DOI:10.1016/j.bse.2010.09.012 |
Zhou RL, Wang HO, Zhao HL. 1999. Response of protective enzymatic system in desert plants grown in different kinds of dunes to atmosphere dehydration and high temperature. Journal of Desert Research, 19(S1): 49-54. |
 2018, Vol. 10
2018, Vol. 10
